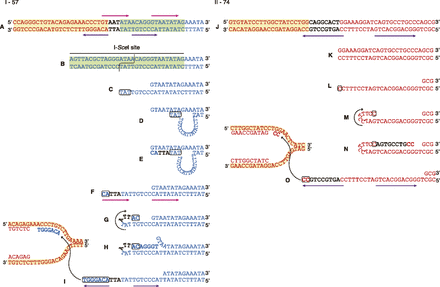
Templated insertions at DEL breakpoints. (A) Sequence of the DEL breakpoint in clone I-57 (Fig. 3A). A net insertion of a 3-bp DNA stretch (5′-AAT-3′, black bold typeface) was found between the 129-derived (red) 5′ breakpoint (chr8: 125087464) and the B6-derived (blue) 3′ breakpoint (I-SceI site inserted at chr8: 125097008) of the DEL interval. Part of the I-SceI recognition site is highlighted in light green. Pink and purple arrows indicate direct and inverted repeats, respectively. (B–I) Multi-step model based on the loop-out and snap-back models proposed in synthesis-dependent microhomology-mediated end joining (Yu and McVey 2010). (B) The I-SceI site on the B6 allele is cleaved by ISceIo transfection. (C) The 5′ end of the DSB site is resected to create a 3′ ssDNA tail (bottom strand). (D) The 3′ ssDNA tail end (5′-TAT-3′, boxed) seeks out its annealing partner on the residual top strand (5′-ATA-3′) by forming a transient looping-out structure and (E) primes strand extension along the template strand, resulting in (F) the formation of short direct repeats (5′-TATATTAC-3′, pink arrows). (G) The extended 3′ tail end (bottom) is then “snapped back” to form a hairpin structure for annealing of the two-base oligonucleotides (5′-AC-3′, boxed) to the complementary sequence (5′-GT-3′) on the same strand. (H) A template-dependent DNA polymerase extends the 3′ end, removing some nucleotides on the top strand. (I) The hairpin structure is unwound, and the short inverted repeats (5′-ACCCTGT-3′, purple arrows) are formed. The newly synthesized 3′ end is then annealed to ssDNA 9.5 kb centromeric to the DSB site via the microhomology (5′-ACAGGGT-3′, blue bold typeface). Interestingly, the annealing partner resides within a genomic inverted repeat (highlighted in yellow) as shown in Figure 5. (J) Sequence of the DEL breakpoint in clone II-74 (Fig. 3B). A net insertion of an 8-bp DNA stretch (5′-CAGGCACT-3′, black bold typeface) is found between the 129-derived 5′ breakpoint (chr8: 124905418) and 3′ breakpoint (chr8: 125257681) of the DEL interval. (K–O) Multistep model: (K) A one-ended DSB is produced 160 kb telomeric to the Aprt locus by an undefined mechanism. (L) The 5′ end of the DSB site is resected to create a 3′ ssDNA tail (bottom strand). (M) The 3′ tail end is snapped back to form a hairpin structure via one-base microhomology (boxed), and (N) primes strand extension along the template strand. (O) The hairpin structure is unwound, and the short inverted repeats (5′-CAGTGCCTGCC-3′, purple arrow) are formed. The newly synthesized 3′ end is then annealed to ssDNA within a genomic repeat (highlighted in yellow), 192 kb centromeric to the Aprt locus via microhomology (5′-CC-3′, boxed).











