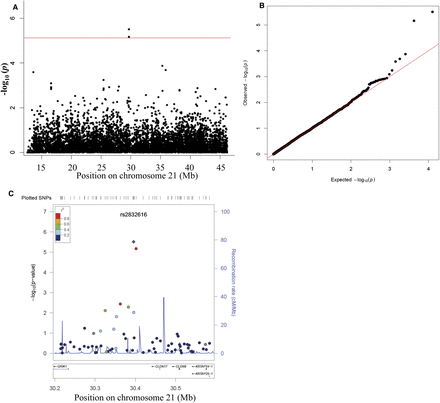
(A) Chromosome 21–wide Manhattan plot and (B) Q–Q plot of SNP genotypic association test P-values for 187 DS-CHD and 151 DS-without CHD using 7238 SNPs on chromosome 21. (Red line) The Bonferroni threshold for chromosome 21–wide α = 0.05. SNPs are plotted in megabases relative to their position on chromosome 21. Two SNPs within the same LD block (r2 = 1) reached chromosome 21–wide significance (P ≤ 0.05) (for details, see Table 1). (C) Regional association plot for the region identified to associate with DS-CHD. The panel shows the recombination rate in the region estimated from HapMap CEU data (http://hapmap.ncbi.nlm.nih.gov/), pairwise LD between SNPs in the region and the SNP identified (purple), and P-values for strength of associations and genes in the region. The r2 values are color-coded according to the scale on the panel.











