
(A,B) The distributions of nucleosome positioning (local dyad positioning score) around representative outron-TSSs (A) and exon-TSSs (B) of expression levels ≥10 and ≥100 (see the distribution around Gaussian TSSs in Supplemental Fig. S2A,B). From these comparisons,
note that the periodic pattern of nucleosome positioning was of substantially greater amplitude for TSSs of greater expression.
(C) Nucleosome positioning (local dyad positioning score) around SL1 sites and 5′ ends of exons. We used all available SL1 sites
without considering their expression levels. (D) Autocorrelation analysis: Let s(x) denote the local dyad positioning score. The autocorrelation defined by 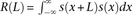 was calculated around representative outron/exon-TSSs, SL1 sites, and 5′ ends of exons, where L is displayed in the x-axis. Sharp peaks at ∼160 bp for outron-TSSs and broad peaks surrounding ∼160 bp for exon-TSSs highlight that arrays of positioned
and phased nucleosomes are more prominent around outron-TSSs than around exon-TSSs. Broad peaks at ∼140 bp are also seen for
SL1 sites and 5′ ends of exons. (E) Pol II occupancy around representative outron/exon-TSSs of expression level ≥10, SL1 sites, and 5′ ends of exons.
was calculated around representative outron/exon-TSSs, SL1 sites, and 5′ ends of exons, where L is displayed in the x-axis. Sharp peaks at ∼160 bp for outron-TSSs and broad peaks surrounding ∼160 bp for exon-TSSs highlight that arrays of positioned
and phased nucleosomes are more prominent around outron-TSSs than around exon-TSSs. Broad peaks at ∼140 bp are also seen for
SL1 sites and 5′ ends of exons. (E) Pol II occupancy around representative outron/exon-TSSs of expression level ≥10, SL1 sites, and 5′ ends of exons.











