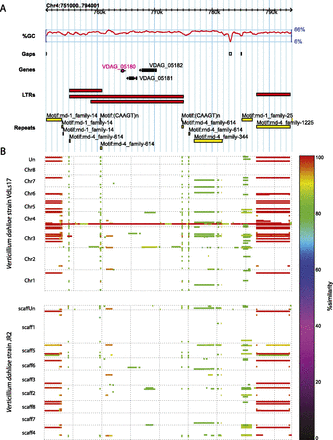
Genomic context of the lineage-specific Verticillium dahliae LysM effector VDAG_05180. (A) Genomic location of VDAG_05180 in V. dahliae strain VdLs17 revealing the presence of flanking gaps, repeats, and a predicted overlapping, but inactive LTR retrotransposon. RepeatModeler families rnd-1_family14, rnd-4_family-614, rnd-1_family-25, and rnd-4_family-1225 are classified as Ty1-Copia, Ty1-Gypsy, TcMar-Fot1, and Ty1-Copia, respectively, by RepeatMasker. (B) Repetitiveness of the genomic context of VDAG_05180 throughout the genomes of VdLs17 (top) and JR2 (bottom). The degree of conservation is indicated by a color scale, showing recent (red) and more ancient (orange to green) multiplications, especially of repeat elements. In addition, the presence of a VDAG_05181 homolog (encoding a tetrahydroxynaphthalene reductase) in the JR2 genome, a duplication of this gene in the VdLs17 genome, and the presence of multiple VDAG_05182 homologs (encoding a Kelch domain-containing protein) in both genomes is shown. The LysM effector gene VDAG_05180 is uniquely found in the VdLs17 genome.











