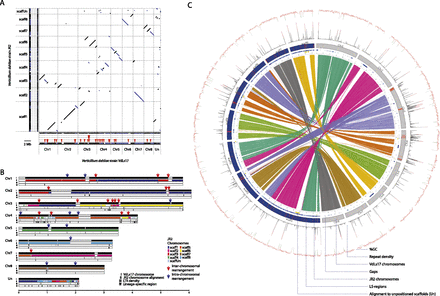
Whole-genome alignment of Verticillium dahliae strains VdLs17 and JR2 reveals extensive chromosomal rearrangements. (A) Whole-genome dot-plot comparison with forward–forward alignments (black) and inversions (blue). Red triangles mark syntenic breakpoints. (Un) Unplaced contigs during optical mapping. (B) Global view of synteny alignments between V. dahliae strains VdLs17 and JR2 and the distribution of LTR retrotransponsons and lineage-specific regions. VdLs17 chromosomes are shown as reference. For each chromosome, row I represents genomic scaffolds (black) on the chromosome separated by scaffold breaks (gray); row II displays syntenic alignment of JR2 chromosomes, highlighting 28 major intra- and interchromosomal rearrangements that are marked by red (inter) and blue (intra) arrows above each breakpoint; row III represents the density of LTR retrotransposable elements calculated in 10-kb windows; and row IV depicts lineage-specific regions that are found in only a subset of strains. (C) Circos diagram illustrating collinear blocks with alignments between VdLs17 (gray) and JR2 (blue) chromosomes, sequence gaps, sequences aligning to unpositioned scaffolds (yellow), lineage-specific sequences (red), repeat density (% coverage of 10-kb window), and GC % (per 10-kb window).











