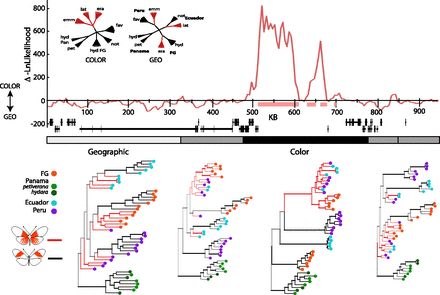
Phylogenetic trees across the D interval. The top panel is the likelihood preference for color-based versus geography-based tree models (inset). Pink bars beneath the plot represent the region where neighbor-joining trees in this sliding window have a monophyletic lineage for the rayed phenotype. These peaks are shown relative to the annotated genes in the interval. The gray bar in the middle represents the five most optimal topological splits used for phylogenetic tree reconstruction, colored by the general history inferred, with black representing a tree clustering by color pattern, light gray a tree clustering by geographic region, and medium gray for transitional topologies. Tree topologies for these divisions are shown below, excluding the tree for the fourth interval, which does not differ substantially from the fifth tree. Nodes supported by a posterior probability >0.95 are represented with bold branches preceding them. Phenotypes are represented by branch color and geographic regions by terminal node color. Internal branches are colored only in cases of clade monophyly of that pattern, otherwise branches are represented in gray. All unrooted tree topologies were arbitrarily rooted at the Panamanian lineage for presentation.











