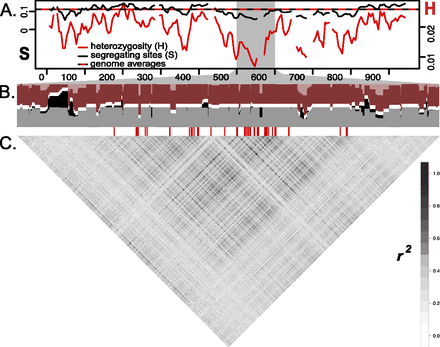
Signatures of selection and recombination across the D interval in the H. erato Peruvian hybrid zone (n = 14). (A) Solid lines show sliding window (15-kb window size, 5-kb step size) values for the number of segregating sites (S, black) and heterozygosity (H, red). Horizontal dashed line represents averages of S and H from genomic regions unlinked to color, with axes scaled to match averages. (B) Haplotypes clustered from 3227 SNPs from the 500–600 kb region, which includes the peak of divergence. Haplotypes from rayed (shaded dark) and postman (shaded light) races were clustered into two groups (K = 2) (colored red and gray). A horizontal white row separates the two clusters. Blocks that have different haplotypes fixed between the two color pattern races are represented by regions with sets of neighboring SNPs in which all rayed individuals are assigned exclusively to the red cluster (red columns shaded entirely dark) and all postman individuals are assigned exclusively to the gray cluster (gray columns shaded entirely light). Blocks where the two color pattern races share haplotypes are represented by regions where individuals of a single color pattern race are assigned to more than one cluster (red columns with light and dark shading; gray columns with light and dark shading). SNPs with fixed allelic differences between color patterns are denoted by red hash marks below the haplotype clustering (data from Fig. 2). (C) Correlation plot of linkage disequilibrium (r2) among 3187 biallelic SNPs across the 500–600-kb window on the D interval.











