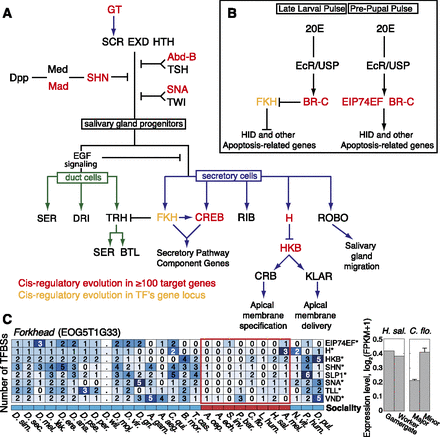
Transcription-factor-mediated signaling pathways controlling salivary gland development. (A) Sex combs reduced (SCR) in combination with extradenticle (EXD) and homothorax (HTH) direct the specification of cells to the salivary gland fate in PS2 of the Drosophila embryo; these TFs are essential for the downstream regulation of genes required for gland cell differentiation and morphogenesis. Boundaries of salivary gland development are restricted along the anterior/posterior axis by abdominal B (Abd-B) and teashirt (TSH), along the dorsal axis by decapentaplegic (DPP) signaling, and along underlying mesoderm by twist (TWI) and snail (SNA). Epidermal growth factor (EGF) signaling determines the decision to differentiate into duct or secretory cells. (B) Regulation of programmed cell death of embryonic salivary gland by 20-hydroxyecdysone (20E). Cell death is inhibited by fork head (FKH) expression in embryonic salivary gland cells. A pulse of 20E at the late larval stage triggers broad (BR-C)–mediated FKH inhibition. A second pulse of 20E at the prepupal stage leads to BR-C and ecdysone-induced protein 74EF (EIP74EF) directed transcription of apoptotic genes, including wrinkled (HID). TFs associated with TFBS evolution in more than 100 target genes (cis-regulatory evolution) (see Fig. 4) (red). The promoter of the fork head locus (yellow), which encodes a TF involved in the regulation of secretory cells, shows significant changes in the gain or loss of promoter TFBSs in ants (trans-regulatory evolution), as detailed in C. (Right) mRNA expression level estimates for fork head, shown for different worker castes in H. saltator (reproductive/non-reproductive) and C. floridanus (major/minor). Error bars indicate standard error over three biological replicates.











