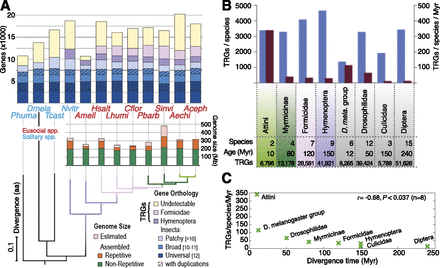
Overview of protein-coding gene composition and genome size in Hymenoptera. (A) Gene and genome content in seven ant species and honeybee (red), with representative solitary insects (blue) as outgroups. Orthology delineation among protein-coding genes from 12 insects identified orthologs present in all (Universal, n = 12) or almost all (Broad, 10 ≤ n ≤ 11) species, conserved as single-copy genes or with paralogs (with duplications). Differential gene losses leave orthologs shared among fewer species across the phylogeny (Patchy, n < 10). Remaining ant genes exhibit orthology with honeybee ([AMELL] Apis mellifera) and/or jewel wasp ([NVITR] Nasonia vitripennis) (Hymenoptera), among ants (Formicidae), or lack orthology (Undetectable). Total estimated genome sizes vary among Hymenoptera, largely due to repetitive regions (orange bars); however, hymenopterans share a nonrepetitive core of ∼200 Mb (green bars). A maximum-likelihood species tree computed from the concatenated alignment of all universal single-copy orthologs confirms the established ant phylogeny (Moreau et al. 2006). Rates of molecular evolution are comparable to the other hymenopterans, flour beetle ([TCAST] Tribolium castaneum; genome size ∼200 Mb), and body louse ([PHUMA] Pediculus humanus; genome size ∼108 Mb), but are much slower than the dipteran representative ([DMELA] Drosophila melanogaster; genome size 175 Mb). The ant species are (HSALT) Harpegnathos saltator; (LHUMI) Linepithema humile; (CFLOR) Camponotus floridanus; (PBARB) Pogonomyrmex barbatus; (SINVI) Solenopsis invicta; (AECHI) Acromyrmex echinatior; and (ACEPH) Atta cephalotes. (B) Occurrence (blue) and emergence rate (red) of taxonomically restricted genes (TRGs) in different taxonomic clades of Hymenoptera (colors) and Diptera (gray). The youngest clades of both Hymenoptera and Diptera exhibit the highest rates of TRG accumulation. Age is measured as the time between the most distant members of each group and hence does not reflect a clade's absolute age. (C) Rate of change of TRGs versus divergence time, for eight species groupings. Pearson's correlation coefficient is shown. P-value was computed using a two-tailed t-test.











