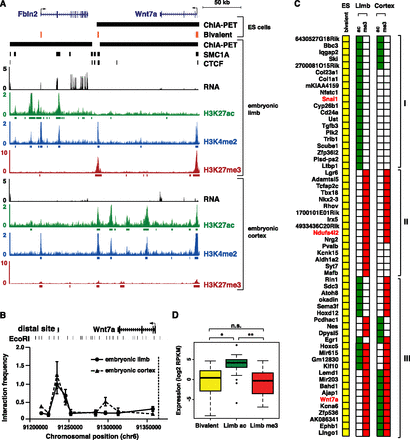
Tissue-dependent activation or repression of poised cohesin interactions. (A) SMC1A/CTCF interaction at Wnt7a in ES cells (Handoko et al. 2011) and limb. Bivalent sites (H3K4me3 and H3K27me3) in ES cells are shown in orange (Mikkelsen et al. 2007). RNA-seq and H3K27ac in E11.5 limb and E14.5 cortex, and H3K27me3 in E11.5 limb and E11.5 cortex are shown (E14.5 cortex RNA obtained from Ayoub et al. (2011). (B) 3C analysis of E11.5 limb and E14.5 cortex interactions between the Wnt7a promoter (dashed line) and distal sites across the locus, normalized to a crosslinking control at Ercc3. Each data point represents average interaction frequency and error bars represent standard error from three independent qPCR reactions (see Supplemental Fig. S5B for biological replicate). (C) Diagram of a subset of bivalent ES cell promoters involved in SMC1A or CTCF ChIA-PET interactions (Handoko et al. 2011). Only interactions with concordant chromatin states at both the promoter and distal site were considered. (I) Interactions resolving to active H3K27ac in limb; (II) interactions resolving to repressive H3K27me3 in limb; (III) interactions resolving to opposite chromatin states in limb and cortex. Gene names in red are involved in interactions detected in both ES cells (Handoko et al. 2011) and limb. The full diagram can be found in Supplemental Figure S5. (D) Gene expression of promoters involved in poised, activated, or repressed interactions. Promoters of H3K27ac marked limb interactions show significantly higher expression than bivalently marked promoters in ES cells ([*] Wilcoxon P-value = 1.6 × 10−11) (Shen et al. 2012) and promoters in H3K27me3 marked interactions ([**] Wilcoxon P-value = 6.1 × 10−10).











