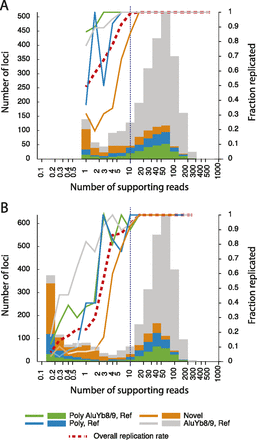
Histograms of the numbers of Alu insertion loci detected in two technical replicates of individual AFP20, binned according to the number of sequencing reads (coverage-corrected to 200,000 indexed, Alu-positive, uniquely mapped reads) supporting them in one replicate, along with the fraction of insertions in each bin that were also observed (at least one supporting read set) in the other replicate. DNA from AFP20 was analyzed twice, once in the ‘African’ library and once in the ‘Variable’ library. (A) Summary of evidence for Alu insertions as detected in the lower-coverage ‘African’ library and their replication rate in the higher-coverage ‘Variable’ library. (B) Summary of evidence for insertions in the ‘Variable’ library and replicated (or not) in the AFP20 sample in the ‘African’ library. In both panels, the histogram bars are sectioned according to the type of Alu insertion observed: green for AluY8/9 insertions that are present in the hg19 human reference genome and previously observed to be polymorphic; blue for Alu insertions present in hg19 and known to be polymorphic but not classified as AluYb8/9; orange for novel Alu insertions detected in the current work, all presumed to be polymorphic; and gray for AluYb8/9 insertions present in hg19 and not known to be polymorphic. The lines indicating replication rates use the same color scheme, with an additional line (red dashed) indicating the overall replication rate. Identical bins and the log-scaled horizontal axis are used in both panels. The vertical dashed line at 10 coverage-corrected reads indicates the threshold used to identify credible insertions.











