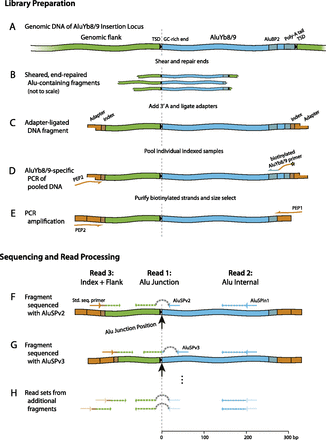
ME-Scan library preparation (A–E) and sequencing (F–H). (A) Genomic DNA (green) containing an AluYb8/9 (blue) in the forward orientation. The 5′ GC-rich region, the 3′ poly-A tail, and the AluYb8/9-specific primer site (AluBP2) are shown as darker segments. The target site duplications (TSD) are indicated by boxed black arrows. (B) DNA samples are fragmented, end-repaired, and an overhanging 3′ A is added. (C) Partially double-stranded oligonucleotide adapters (orange) with 9-bp indexes (darker segment) are ligated onto fragment ends. Indexed samples are then pooled. (D) AluYb8/9 element junctions are targeted by PCR using a biotinylated AluYb8/9-specific primer (AluBP2) and adapter primer PEP2. Biotinylated DNA molecules are then purified using streptavidin-coated paramagnetic beads. (E) Reamplification of the library with primers PEP1 and PEP2 (orange arrows). (F–H) The pooled junction library is then sequenced with a three-read design. Primers are shown as arrows, sequencing reads as dashed lines. The first sequencing read (50 nt) extends from AluSPv2 (F) or AluSPv3 (G). To skip the 5′ end of the Alu insertions, which would be identical over the entire flow cell and therefore difficult for Illumina's software to process, the first 30 cycles of nucleotide synthesis are carried out without collecting data (represented by a dashed gray ‘hop’). The second read proceeds for 57 bp from AluSPIn1. The third read is generated using Illumina's standard second-end read primer and consists of a 9-bp index, a ‘T,’ and 36 bp of genomic sequence from a fragmentation site 50–300 bp upstream of the Alu insertion. Read sets are generated from multiple different fragments representing each AluYb8/9 insertion in the library (H). Each insertion is uniquely identified by its “Alu Junction Position” (dashed line and large arrow in F and G).











