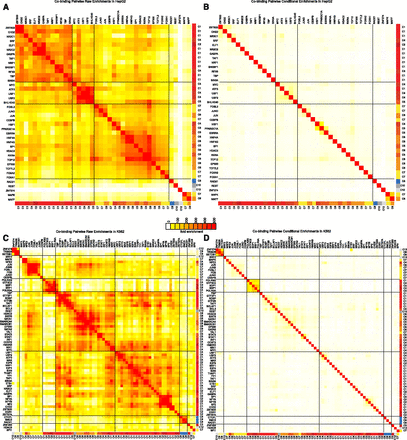
Pairwise regulator cobinding enrichments are captured by chromatin state preferences. (A) Pairwise regulator cobinding enrichment for all pairs of regulators in HepG2 show strong groups of cobinding. The regulators in each group are typically assigned to the same set of chromatin state cluster preferences from Figure 1. Enrichment levels up to 500-fold are found for the full pairwise enrichment table. Rows of the table have been ordered to maximize correlation of neighboring rows. Black lines correspond to groups of highly enriched pairs of regulators that emerge from this ordering. (B) After conditioning on the chromatin state preferences for each regulator (see Methods), the pairwise regulator enrichments are dramatically reduced. (C,D) The same as A and B except for K562. Other cell types can be found in Supplemental Figure 21.











