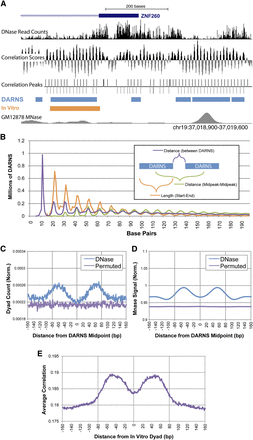
Hidden Markov model (HMM) identified DNase annotated regions of nucleosome stability (DARNS). (A) UCSC Browser screen shot of DNase-seq read counts, correlation scores, correlation peaks, and HMM identified DARNS (light blue boxes). In vitro positioned nucleosomes (orange box) (Valouev et al. 2011) and the in vivo GM12878 MNase-seq show a similar overlap with DARNS. (B) Length of and distance between DARNS display an ∼10-bp pattern. (C) Distribution of in vitro dyads shows twin crests of significant enrichment (∼[−80:−45] and [45:80]) around DARNS midpoint. (D) Distribution of in vivo GM12878 MNase signal shows twin crests of significant enrichment (∼[−80:−25] and [25:80]) around DARNS midpoint. (E) Distribution of average correlation at peaks from both strands shows a dip in scores near in vitro dyads.











