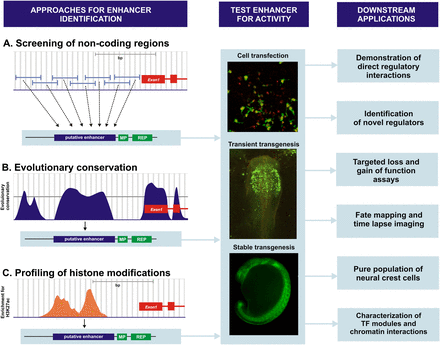
Strategies for cis-regulatory analysis of the neural crest. (A) Early approaches for identifying enhancers of neural crest genes included screening long stretches of noncoding DNA. Fragments from the locus of the gene of interest were cloned upstream of a minimal promoter and a reporter gene and tested in vivo by cell transfection or transgenesis. (B) Sequencing of vertebrate genomes facilitated the search for cis-regulatory modules. Computational comparison between different species reveals evolutionary conserved regions (ECRs) that are putative regulators of nearby genes. The ECRs are subsequently tested for activity by stable or transient transgenesis in different model organisms. (C) Novel approaches allow for genome-wide analysis of the cis-regulatory modules. Profiling of histone modification through ChIP-seq allows mapping of chromatin mark patterns and identification of active and poised enhancers. This method was used to annotate enhancers that are active in human neural crest cells induced from human embryonic stem cells (Rada-Iglesias et al. 2012). Once enhancers are validated in vivo, they become valuable tools that can be exploited in different contexts. (MP) Minimal promoter; (REP) reporter gene; (TF) transcription factor.











