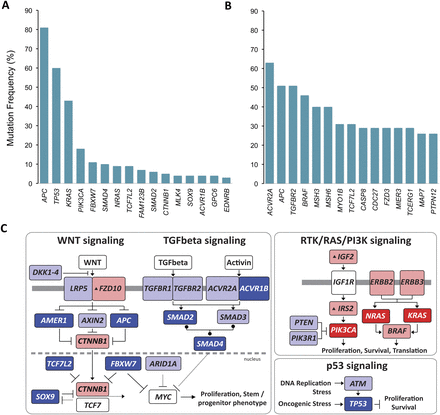
Significantly mutated genes and principal cancer pathways deregulated by somatic mutations in human colorectal carcinoma. Patients are divided into two groups based on mutation rate. All genes shown are significantly mutated with a false discovery rate of less than 0.1. (A) Profile determined from 193 patients with chromosome instable, low mutation rate, disease (see Fig. 2A, CRC MSS). (B) Profile determined from 29 microsatellite instable CRC plus 7 POLE-mutated patients (see Fig. 2A, CRC MSI and CRC MSS POLE). (C) Principal cancer pathways deregulated by somatic mutation in CRC. Alterations are defined by somatic mutations, homozygous deletions, high-level focal amplifications, and, in some cases, by significant up- or down-regulation of gene expression (black up-triangle). All genes from Figure 3 except MLK4, GPC6, and EDNRB can be placed in one of the four pathways shown here. WNT signaling is disrupted by one or more mutations in 93% of patients; TGFbeta signaling is disrupted in 26% of all patients with a low mutation rate and in 94% of patients, and RTK/RAS/PI3K signaling is disrupted in over 80% of patients. (Red) Activated genes; (blue) inactivated genes. Deep red or blue are genes on the significantly mutated list from panels A and B. Lighter shaded genes are not mutated significantly in this cohort but contribute to pathway disruption in some patients. Panels A and B adapted from Figure 1, and panel C from Figure 4, of The Cancer Genome Atlas Research Network (2012a).











