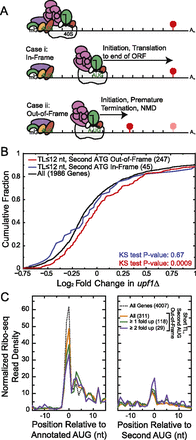
Short TL genes are enriched for nonsense-mediated mRNA decay (NMD) targets. (A) Model predicting why short TL genes with a second out-of-frame AUG are NMD targets (not to scale). Failure to identify the cap-proximal AUG in short TLs results in scanning and recognition of a second, downstream AUG. If the second AUG is out-of-frame, it results in premature termination and NMD. For simplicity, the eIF4F complex is drawn on the cap during scanning, though some or all of its subunits may remain associated with the small ribosomal subunit (Jackson et al. 2010; Aitken and Lorsch 2012). (B) Fold change in steady-state mRNA levels for short TL genes in upf1Δ cells. Genes with short TLs exhibit a significant shift toward increased RNA levels only when the next AUG encountered is out-of-frame. The number of genes in each group is indicated in parentheses. (C) Ribosome density analysis of genes with a second AUG out-of-frame, using Ribo-seq from glucose-starved yeast. The dotted line shows all genes; solid lines show short TL genes with second AUG out-of-frame. Fold up indicates genes whose mRNAs are increased in the upf1Δ microarray data.











