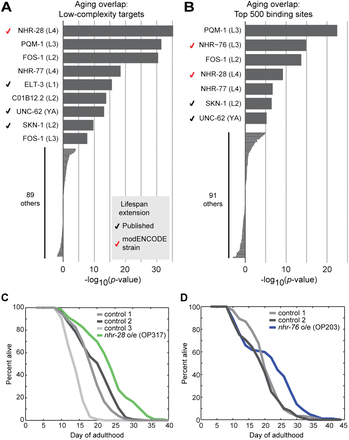
Identification of candidate regulators of aging. (A,B) The ChIP-seq–based screening approach described in Figure 5 was applied to the set of genes with altered expression during aging (Budovskaya et al. 2008). For each ChIP-seq data set, the correlation was calculated twice: first, using all low-complexity binding sites (A), and second, using the 500 binding sites with the highest naïve Bayes–derived score as described in Figure 5D (B). (Bars) The P-value (log10) between age-regulated genes and the indicated ChIP-seq targets, with enrichment indicated by positive values and depletion indicated by negative values. For nine transcription factors, significant overlap (P < 10−5) was observed using at least one of the two approaches; six of nine were significant with both. (Black checkmarks) Three factors (ELT-3, UNC-62, and SKN-1) for which modulation has been shown to increase lifespan (Curran and Ruvkun 2007; Budovskaya et al. 2008; Tullet et al. 2008). (Red checkmarks) Factors for which the modENCODE strain (which contains an integrated multi-copy fosmid containing the listed transcription factor) has an extended lifespan. (C,D) Lifespan of modENCODE-generated strains in which a strain containing a fosmid with C-terminal GFP-tagged nhr-28 or nhr-76 was compared with controls. Days of adulthood are indicated on the x-axis, and the percentage of worms remaining alive is indicated on the y-axis. (C) Strain OP371 (containing a fosmid with GFP-tagged nhr-28) was compared with three controls (RW10780, RW11206, and RW11175). The strain overexpressing nhr-28 shows 15%–30% extension of lifespan relative to the various controls (all P < 10−5 by log-rank test). (D) Strain OP203 (containing a fosmid with GFP-tagged nhr-76) showed a 7%–15% increase in mean lifespan relative to two controls (RW10780 and RW11206) (P < 0.01 against either). Lifespan data shown are from strains that were backcrossed twice to wild-type; each lifespan experiment was performed twice before backcrossing and gave similar results (Supplemental Table 5).











