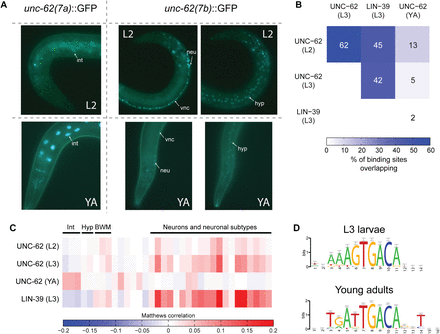
Distinct sets of targets bound by UNC-62 in L2/L3 larvae and adults. (A) Strains expressing isoform-specific unc-62 translational reporters show stage- and tissue-specific expression (Van Nostrand et al. 2013). (Left) unc-62(7a):GFP is not observed in early larval stages but is highly expressed in the intestine in late larval stages and young adults (YA). (Right) unc-62(7b):GFP is not observed in the intestine at any stage but is expressed in the hypodermis (hyp), the ventral nerve cord (vnc), and other neurons (neu). Strains were imaged in a glo-4(ok623) background to limit gut autofluorescence. (B) The overlap of ChIP-seq binding sites for UNC-62 in L2 and L3 larvae, young adults (YA), and HOX transcription factor LIN-39 in L3 are shown as the percentage of binding sites in the smaller set that are also bound in the larger. (C) Targets bound by UNC-62 in L2, L3, and YA, as well as those bound by LIN-39 in L3 larvae, were compared to genes enriched for expression in various tissues (Roy et al. 2002; Zhang et al. 2002; Colosimo et al. 2004; Fox et al. 2005; Pauli et al. 2006; Von Stetina et al. 2007; Spencer et al. 2011). Colors indicate the correlation between low-complexity target genes and genes with tissue-enriched expression for the indicated tissue. Tissues are clustered according to broad tissue types, and the specific tissue for each column is listed in Supplemental Figure 2 and Supplemental Table 3. (Int) Intestine; (Hyp) hypodermis; (BWM) body wall muscle. (D) A de novo motif search with RSAT (Thomas-Chollier et al. 2012) identifies sequences significantly enriched in the 100-bp central core region of UNC-62 binding sites. A total of 200-bp flanking regions on either side of this core were used as the background sequence set. Both motifs contain the D. melonogaster Homothorax motif (TGACA) (Noyes et al. 2008).











