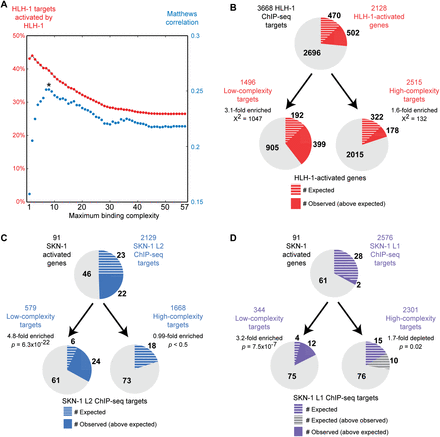
Low binding site complexity correlates with factor-responsive expression. (A) Using 4191 significant HLH-1 binding sites identified by the modENCODE Consortium (Niu et al. 2011), the set of genes with HLH-1 binding sites with complexity less than or equal to n (for n = 1–57) was identified. Each set was then compared to 2128 genes activated upon HLH-1 overexpression (Fukushige et al. 2006; Fox et al. 2007), with the percentage of directly bound targets activated indicated in red. Except for complexities of two or less, binding sites with lower complexity had higher precision in predicting HLH-1–activated genes. By use of the Matthews correlation coefficient (in blue) to weight both false-positives and false-negatives, a complexity of eight or less was identified as optimal for predicting factor-responsive targets (indicated by *). (B) Using only binding sites with complexity of eight or less significantly improves prediction of HLH-1–responsive binding. (Circles) The set of genes with any HLH-1 binding site (top) or only those with low-complexity or intermediate/high-complexity binding sites (bottom). The overlap with HLH-1–activated genes is indicated in red, with the expected overlap indicated by white hash marks. Significance of enrichment was calculated by Yates' χ2 test. (C,D) The same complexity criteria significantly delineate SKN-1 targets in L2 and L1 larvae. Circles indicate 91 SKN-1–activated genes (Park et al. 2009), with the percentage overlap with SKN-1 ChIP-seq data sets indicated in blue and expected overlap indicated by white hash marks. Low-complexity regions for SKN-1 in L2 larvae (C), and L1 larvae (D), are enriched for genes responsive to skn-1 knockdown. Enrichment significance was determined by Fisher's exact test. The set of all SKN-1 targets in L1 was not significantly enriched for SKN-1–activated genes (1.1-fold-enriched, P > 0.5), indicating that the correlation between SKN-1 binding in L1 larvae and SKN-1–responsive expression is only observed when binding site complexity is taken into account.











