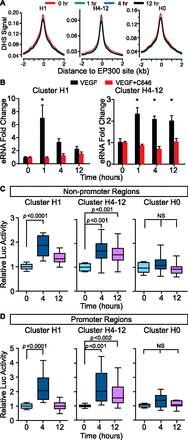
Dynamic H3K27ac regions had functional properties of transcriptional regulatory regions. (A) DNase-seq showed that dynamic, EP300-associated sites are hypersensitive to DNase I digestion, but the sensitivity did not change substantially during the VEGFA stimulation time course. DNase signal is plotted as reads per 10−6 mapped reads. (B) eRNA expression from dynamic H3K27ac regions with or without EP300 inhibition by C646. Each bar represents the average of 5–6 different regions (individual data are shown in Supplemental Fig. 7). (*) P < 0.05. (C,D) Luciferase activity of dynamic H3K27ac regions at promoter and nonpromoter sites. Luciferase activity was expressed relative to the activity measured at time 0. n = 5–8 per group. Line, bar, and whiskers represent median, quartiles, and min-max values, respectively.











