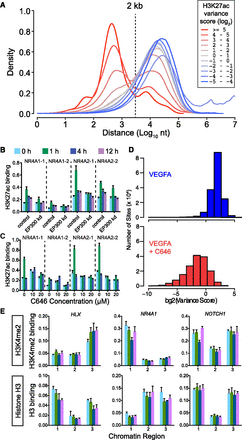
Dynamic H3K27ac regions were associated with EP300. (A) Distance relationship of H3K27ac sites to nearest EP300-occupied region as a function of H3K27ac variance score. The H3K27ac regions with the highest variance score were predominantly located within 2 kb of EP300. (B,C) EP300 is required for dynamic H3K27 acetylation. ChIP-qPCR of H3K27ac in HUVEC cells treated with VEGFA and EP300 siRNA (B) or small molecule EP300 acetyltransferase inhibitor C646 (C). Two dynamic H3K27ac regions near each of the indicated four genes were tested (see also Supplemental Fig. 5A,B). (D) C646 pretreatment suppresses H3K27ac change at highly variant sites. We considered genomic windows with a log2 variance score of at least 2 and at least the sum of 10 reads across all four time points. We recalculated the variance score for each site under VEGFA stimulation in the absence or presence of C646 for hours 0, 1, and 4. The histogram shows that C646 reduced the variance score at the vast majority of sites. (E) Chip-qPCR of H3K4me2 and total histone H3 binding at three dynamic H3K27ac sites showed that altered nucleosome positioning is unlikely to account for VEGFA-stimulated changes in H3K27ac. Numbered chromatin regions are indicated in Fig. 1D.











