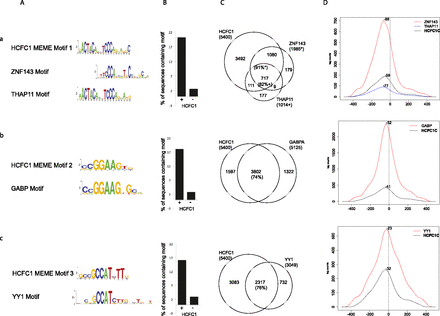
Identification of transcription factors associated with HCFC1-bound TSSs. (A) The HCFC1 MEME Motif logo identified in HCFC1-bound TSS-associated sequences is displayed above the most similar TRANSFAC motif(s). For HCFC1 MEME Motif 1, the motif corresponding to the experimentally described sequence for mouse Thap11 (Ronin) is also shown (Dejosez et al. 2010). (a) ZNF143 and THAP11; (b) GABP; (c) YY1. (B) Percentage of HCFC1-bound (+) or -unbound (−) sequences that contain the HCFC1 MEME motif. (C) Venn diagrams showing the overlap between HCFC1-bound TSS regions and ZNF143, THAP11, GABPA, and YY1-bound TSS regions as identified by ChIP-seq. The percentages of TSS regions that contain the transcription factor-binding sites and are HCFC1 positive are given. (*) Percentage of ZNF143 binding sites that are HCFC1 positive; (+) percentage of THAP11 binding sites that are HCFC1 positive. (D) Cumulative mapping of sequence tags around −250 to +250 of TSSs (determined as in Fig. 1E) shared between HCFC1C and the corresponding transcription factor. The most enriched position is indicated as the distance in base pairs from the TSS, which is indicated as a dashed vertical line.











