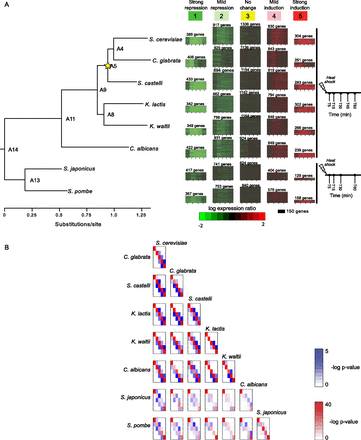
Heat shock transcriptional response in eight Ascomycota species is captured by five modules. (A) Expression modules identified by Arboretum in the transcriptional response to heat shock in eight species. Shown are the expression modules (1–5, heat maps, middle) in each of eight species (species tree, left) at denoted time points prior to and following heat shock (time axis, right). Color bar denotes expression relative to pre-stress time zero. (Red) induced; (green) repressed; (black) no change. Each heatmap shows the expression profile of all genes assigned to that module in a given species. The heatmap height is proportional to the number of genes in the module (marked on top). All modules in one column are mapped to the same ancestral module ID (1–5, top) at the LCA of these eight species. (B) Overlap of modules between species. Shown is the degree of overlap in orthologous genes between every pair of modules 1–5 (rows and columns in each matrix) in every pair of extant species. Diagonal elements (red): overlap between modules of the same ID; off-diagonal elements (blue): overlap between modules of different IDs. Red and blue intensity is proportional to −log (P-value) of the hypergeometric distribution (color scales, right).











