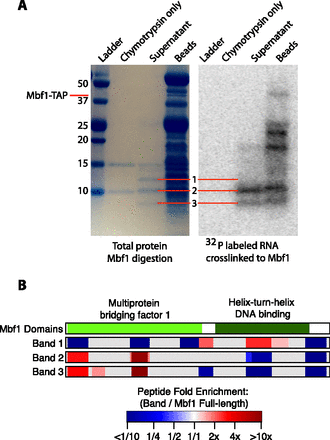
Mapping the RNA-binding domain of Mbf1. (A) SDS-PAGE analysis of protein fragments resulting from partial digestion of Mbf1 with the protease chymotrypsin. The lanes from left to right contain the ladder, chymotrypsin only, the supernatant of protein fragments liberated by chymotrypsin digestion of Mbf1, and the protein fragments remaining on the beads. A total protein stain is shown on the left, and the radioactive image of the same gel is shown on the right. Gel images were scaled and aligned to facilitate direct comparison of the visible bands. The radioactive image shows a signal from 32P-labeled RNA fragments cross-linked to Mbf1. (B) A diagram showing the domains of Mbf1 and the position and enrichment relative to full-length Mbf1 for all the peptides that were detected in each sample. The fold enrichment of the normalized intensity of each peptide relative to undigested Mbf1 for each of the three bands is represented by a color gradient ranging from dark blue for less than one-tenth fold enriched, to white for no enrichment, to dark red for greater than 10-fold enriched. (Gray) Areas of the protein for which no peptides were detected.











