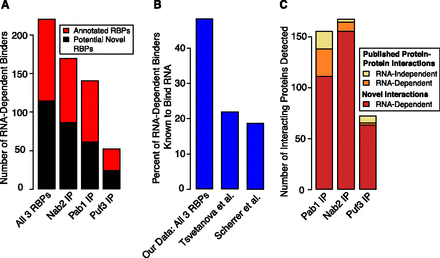
Barplots showing enrichment of known RNA-binding proteins and the RNA dependence of published protein–protein interactions. (A) The fraction of proteins we identified as RNA-dependent binders that are known RBPs (defined as those that have domains known to bind RNA or have a molecular function of RNA binding in the Gene Ontology database). Known RNA-binding proteins are significantly enriched among the proteins interacting in an RNA-dependent manner with Pab1, Nab2, and Puf3 (for example, P-value of 2 × 10−35 for the union of all three data sets). (B) The percentage of proteins known to bind RNA in the set of 220 RNA-dependent binders in our combined data set, compared with the top 220 proteins from the protein array data from two previously published attempts to identify proteome-wide RNA–protein interactions. All had significant enrichment of known RBPs, but the enrichment seen with the set of proteins identified by our method was much greater (hypergeometric P-values 2 × 10−35, 2 × 10−4, and 2 × 10−5 for this study, Scherrer et al. 2010, and Tsvetanova et al. 2010, respectively). (C) A barplot showing the RNA dependence of published high-confidence protein–protein interactions and also the number of novel interactions (RNA dependent) we observed with Pab1, Nab2, and Puf3.











