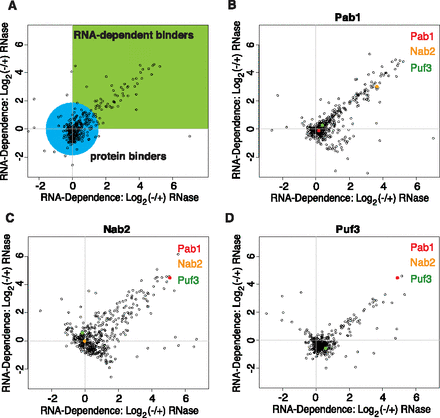
Overview of RNA-dependent interaction data. A scatterplot of the RNA-dependent enrichment values as the log base 2 (−/+) RNase ratios for two replicates (with inverted labeling schemes) of each protein that we purified. The reproducible RNA-dependent binders form a tail in quadrant 1. (A) A plot of example data with a green box indicating the quadrant where RNA-dependent binders are expected to be found and a blue circle indicating where RNA-independent binders are expected to be found. These colored regions are broad generalizations only and were not used for actual data analysis. (B) A scatterplot of the log base 2 (−/+) RNase ratios for the replicate experiments with Pab1. The points representing the proteins Pab1, Nab2, and Puf3 are highlighted in red, yellow, and green, respectively. (C) The same scatterplot for experiments with the protein Nab2. (D) The same scatterplot for experiments with the protein Puf3.











