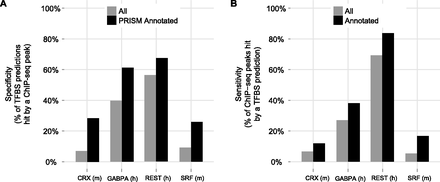
Overlap of PRISM binding site predictions with ChIP-seq. PRISM binding site predictions for CRX (m = mouse), GABPA (h = human), REST (human), and SRF (mouse) were overlapped with ChIP-seq binding sites for the same four factors. Overlap was observed both for the full set of binding site predictions and a subset annotated by PRISM in a particular functional role (“sensory perception of light stimulus” for CRX, “translation” for GABPA, “neurotransmitter transport” for REST, and “actin cytoskeleton” for SRF). (A) Percent of PRISM binding site predictions hit by a ChIP-seq peak. (B) Percent of ChIP-seq peaks hit by a PRISM binding site prediction. Only ChIP-seq peaks with a match to the transcription factor motif in the reference species were considered (CRX: 74%; GABPA: 81%; REST: 60%; SRF: 34%).











