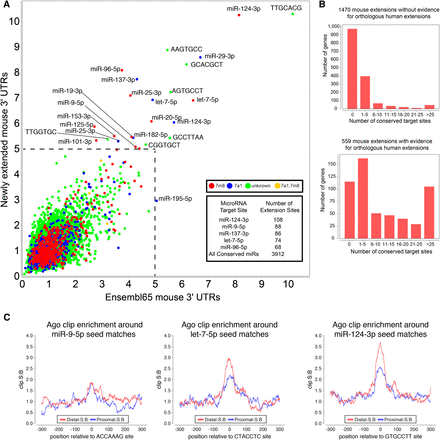
Novel 3′ UTR extensions harbor thousands of functional miRNA target sites. (A) Signal-to-background ratio (S:B) of 7-mers found in the proximal 3′ UTR annotations compared with the novel extended 3′ UTR region annotated in mouse from all tissues analyzed. Note that target sites for several well-characterized neural miRNAs are found among the most well-conserved 7mers in both proximal and novel extended 3′ UTR regions, including miR-124, miR-137, miR-9, let-7, miR-96, and miR-125. Supplemental Figure S10 demonstrates that the signal for neural miRNA seed matches is driven by genes with neural-expressed 3′ UTR extensions. (B) Analysis of seed matches to mammalian-conserved miRNAs, that are present among mouse 3′ UTR extensions that lack companion expression evidence for an orthologous 3′ UTR extension in human (top graph) or that do have such experimental evidence for a human extension (bottom graph). The proportion of conserved miRNA binding sites is much higher among genes with evidence for a conserved 3′ UTR extension. (C) Regions surrounding miRNA target sites located in proximal (in blue) and novel distal 3′ UTR mouse extensions (in red) show enrichment of Ago HITS-CLIP tags over background. The signal:background (S:B) of clip tags at let-7 and miR-124 seed matches is actually higher in the novel 3′ UTR extension regions.











