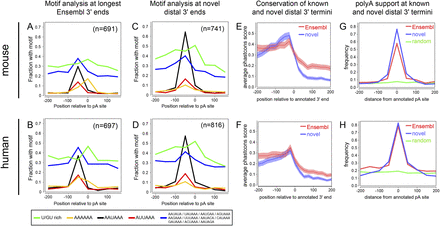
Sequence and conservation features of known and novel 3′ termini. All analyses in this figure concern those mouse (top graphs) and human (bottom graphs) genes whose 3′ UTRs were confidently extended in this study. These comprise 691 Ensembl65 mouse gene models for which we precisely annotate 741 novel 3′ termini in one or more tissues, and 697 Ensembl65 human genes for which we precisely annotate 816 novel 3′ termini in one or more tissues. (A,B) Motif frequency in 50-nt bins in the vicinity of annotated 3′ termini. Motifs are listed at bottom, and include the downstream U/GU-rich region that promotes 3′ cleavage, the canonical PAS AAUAAA and its most common variant AUUAAA, a panel of low-frequency PAS variants, and genomically encoded hexa-A tracts. As expected for annotated mouse (A) and human (B) 3′ termini, there is strong positional enrichment of functional PAS upstream of the polyadenylation site and U/GU downstream. The collection of low-frequency PAS variants exhibits a broad background frequency, with mild enrichment at the normal location of canonical PAS. Unexpectedly, we observed enrichment of A6 at annotated 3′ termini, potentially reflecting internal priming events in this collection of curated 3′ termini. (C,D) The frequency and positional specificity of PAS and U/GU motifs in our novel mouse (C) and human (D) 3′ termini are relatively similar to known termini but lack substantial A6 enrichment at transcript ends. (E,F) Analysis of average phastCons scores in the vicinity of known and newly annotated 3′ termini in mouse (E) and human (F) shows that both populations of termini exhibit selective constraint that rises to a peak in the local sequence upstream of 3′ termini, and drops sharply in the downstream sequence. Note also that the aggregate conservation of the last ∼500 nt of proximal 3′ UTR sequences is higher than that of the distal novel 3′ UTR sequences, but the overall level of conservation 3′ of our mouse and human extensions drops to background. (G,H) Analysis of location of polyA-seq tags relative to known and newly annotated 3′ termini shows a similar positional enrichment at transcript 3′ termini. Comparison with a randomly selected set of 3′ ends from these transcripts shows no positional enrichment of polyA-seq tags, indicating that our novel annotations include genuine 3′ ends.











