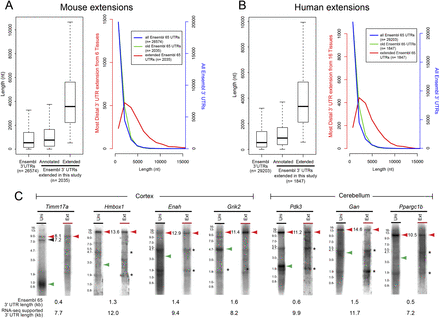
Reevaluation of mouse and human RNA-seq data reveals abundant 3′ UTR extensions. (A, left) Box plot comparing aggregate Ensembl v65 3′ UTR lengths (longest annotation per terminal exon) and those Ensembl v65 3′ UTRs that were specifically extended in this study. (Right) Histogram of the same data to highlight the abundance of newly annotated long 3′ UTRs. (B) Analyses similar to A, except plotting known and novel Ensembl v65 human 3′ UTRs. (C) Northern analysis validates the stable accumulation of many transcripts utilizing very distal polyadenylation signals in cerebellum or cortex, in several cases yielding 3′ UTRs >10 kb in length. Green arrowheads indicate predicted mRNA length of Ensembl v65 gene model. Red arrowheads indicate inferred mRNA lengths of novel 3′ UTR extension isoforms. Asterisks denote background hybridization to ribosomal RNAs.











