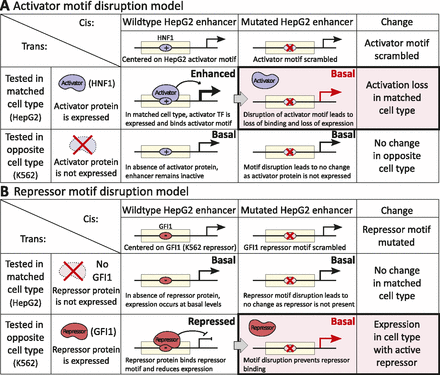
Enhancer activator and repressor models. In our model of enhancer activity, the cell-type specificity of enhancers is maintained by the combined action of activators (such as HNF1 and HNF4 for HepG2 enhancers) that are expressed and bind in the matched cell type (HepG2), and the action of repressors (such as GFI1) that are expressed and bind in the unmatched cell type (K562). (A) Predicted enhancer activators are expressed in the cell type of enhancer activity, and their motifs are enriched within active enhancers. Disruption of the predicted activator motif leads to reduced reporter expression as the activator no longer binds its target motifs. (B) Predicted enhancer repressors are expressed in the other cell type and serve to reduce expression of the reporter gene, by preventing activator binding in the enhancer region or neighboring promoter. Disruption of the repressor motifs shows an effect only in the unmatched cell type, where binding of the repressors is disrupted, thus leading to derepression.











