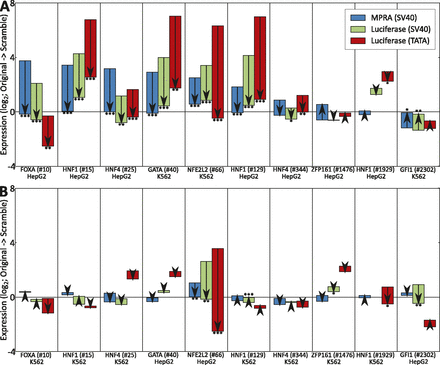
Robustness to tested sequence length and promoter type. (A) Comparison of MPRA (blue) vs. luciferase reporter assays (green/red) using 500-bp sequences instead of 145-bp and alternate promoters. For each of the 10 candidate enhancers, we list the predicted regulator, the enhancer ID (Supplemental Data S1), and the cell type in which the element was tested (matched cell type for predicted activators, unmatched for predicted repressors). Each bar indicates the expression of the original sequence and the effect of motif scrambling (direction of the arrow). MPRA experiments used 145-bp sequences centered on the motifs and a strong SV40 promoter (blue), and luciferase experiments used 500-bp sequences centered on the motifs with either a strong SV40 promoter (green) or TATA promoter (red). Data is normalized by subtracting from each expression value the mean for scrambles in that cell line across these 10 sequences. Asterisks indicate significance values using a t-test on the individual replicate values for the sequences (*) P < 0.05, (**) P < 0.01, (***) P < 0.001; see Methods (Mann-Whitney P-values are available in Supplemental Table S5). (B) Results for each of the sequences tested in A for the reverse cell type where the factor was not predicted to be active. A significant and large change was seen for NFE2L2 (#66), consistent with MPRA results. In addition, we observe significant, albeit smaller, luciferase changes for HNF1 (#129), ZFP161 (#1476), HNF1 (#1929), and GFI1 (#2302). Luciferase SV40 values for HNF1 (#1929) in K562 are absent due to a sample tracking error (see Supplemental Table S5).











