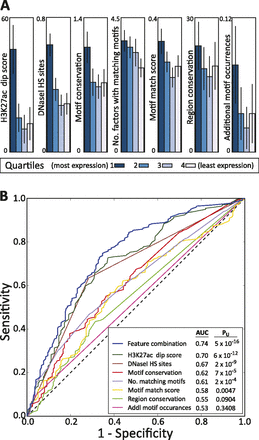
Importance of sequence context for enhancer function. (A) Association of top scoring enhancers with: the average H3K27ac signal value in the matched cell type 200 bp away, minus the value centered on the motif (in 25-bp windows); overlap with DNase I annotations in the matched data (Song et al. 2011); the raw motif conservation score (Kheradpour et al. 2007; Lindblad-Toh et al. 2011); the number of factors with matching motifs in regions outside of the motif match in the tested sequence; the strength of the motif match; the number of bases indicated as conserved by SiPhy-ω 12-mers (Garber et al. 2009); and the number of matches to the tested motif within the tested sequence. (B) Predictive power for recognizing enhancers that are likely to show high wild-type reporter expression based on each of these individual features and a combination of features using logistic regression (Hall et al. 2009).











