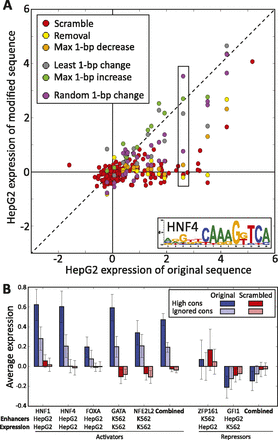
Summary of motif manipulation results for all activators and repressors tested. (A) Average reporter gene expression for 160 predicted HepG2 enhancers centered on conserved HNF4 motifs for wild-type construct expression (x-axis) and modified construct expression (y-axis) for different modifications. A total of 160 constructs with scrambled motifs (red) consistently lie near the y-axis (no reporter expression), confirming the necessity of the conserved HNF4 motif. Five additional motif modifications were tested for the 15 most conserved HNF4 motifs. The preponderance of disruptive modifications (red, yellow, and orange points) showing decreased reporter expression (below the diagonal) demonstrate the dramatic reduction of enhancer activity for the most disruptive mutations, while the presence of neutral (gray) or motif-strengthening (green) modifications near and above the diagonal highlight the specificity of mutations to those that disrupt recognition of the motif. Box indicates example shown in Figure 2A–C. (B) Comparison of reporter expression for enhancers centered on five activators in the matched cell type and two repressors in the unmatched cell type. For the five predicted activators, wild-type reporter expression is higher for 160 enhancers centered on conserved motifs (dark blue) than for 160 enhancers centered on motifs ignoring conservation (light blue), and it is significantly reduced after motif scrambling (red, pink). For the two predicted repressors, motif scrambling results in increased reporter expression in the unmatched cell type (see model in Fig. 6). Error bars represent 95% confidence interval on the mean. Additional bar plots in Supplemental Figure S4. All statistics are shown in Supplemental Figure S2. All expression values in this figure are computed as described in the Methods.











