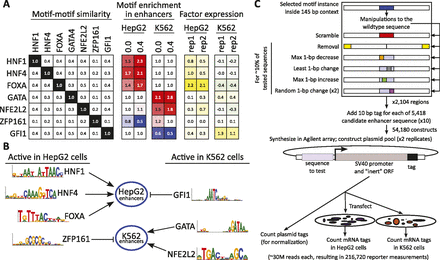
Selection of activator and repressor motifs. (A) Predicted activator and repressor motifs were chosen based on their lack of similarity to each other (left) (Supplemental Fig. S2); fold-enrichment for activators (red) and fold-depletion for repressors (blue) in the cell line of interest (middle); and microarray expression (Ernst et al. 2011) of the corresponding factor in the target cell line (log2, right). Black-white, red-blue, and green-yellow color gradients are used for emphasis, but all values are indicated. (B) Predicted activators and repressors for each cell type and corresponding motifs. HNF1, HNF4, and FOXA are predicted to act as activators of HepG2 enhancers in HepG2 cells. GATA and NFE2L2 are predicted to act as activators of K562 enhancers in K562 cells. GFI1 is predicted to act as a repressor of HepG2 enhancers in K562 cells, and ZFP161 is predicted to act as a repressor of K562 enhancers in HepG2 cells. Details on selection criteria and motif sources are available in Supplemental Figure S2. (C) For each of 2104 predicted enhancer regions, we designed between two and eight variants (colors as in Fig. 3A), each tested in two biological replicates in two cell lines, using 10 different tags per sequence. We also sequenced the plasmid library directly to provide tag counts used for normalization. A single Agilent array is thus used to obtain 54,180 reporter expression levels for 5418 enhancer variants.











