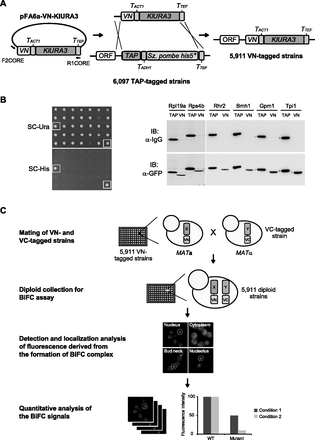
Construction and utilization of the VN fusion library. (A) Construction of the VN fusion library. A DNA fragment containing the VN tag and KlURA3 marker sequences was amplified by PCR, using the pFA6a-VN-KlURA3 vector as the template and the primers F2CORE and R1CORE, and substituted for the C-terminal TAP tag sequence in the chromosome of each strain of the TAP fusion library by homologous recombination. KlURA3, TACT1, TADH1, and TTEF represent Kluyveromyces lactis URA3, S. cerevisiae ACT1 terminator, S. cerevisiae ADH1 terminator, and Ashbya gossypii TEF terminator, respectively. (B) Confirmation of switching from the TAP tag to the VN tag. Transformed cells obtained on SC-Ura plates were replica-plated onto SC-His plates and incubated at 30°C for 3 d (left panel). The cells selected on SC-Ura medium and counter-selected on SC-His medium were regarded as candidates for harboring correctly switched epitopes. The cells that failed in counter-selection by SC-His are represented by open squares. Western blot analysis was performed to confirm correct switching of the VN tag from the TAP tag (right panel). Six highly abundant proteins (Rpl19a, Rps4b, Rhr2, Bmh1, Gpm1, and Tpi1) were selected for Western blot analysis. Both the host cells expressing the corresponding C-terminally TAP-tagged proteins (MK0074, MK0076, MK0078, MK0080, MK0082, and MK0084) and the VN-switched cells (MK0075, MK0077, MK0079, MK0081, MK0083, and MK0085) were grown to mid-log phase in YPD medium at 30°C. Total proteins were extracted, and immunoblotting was performed with HRP-conjugated anti-mouse IgG and anti-GFP antibodies. (TAP) Host strain carrying the corresponding C-terminally TAP-tagged protein. (VN) Epitope-switched strain carrying the corresponding C-terminally VN-tagged protein. (C) Schematic diagram of the genome-wide analysis of in vivo PPIs with the VN fusion library. For the genome-wide BiFC analysis, each strain of the VN fusion library is mated with a MATα strain expressing a protein of interest tagged with the C-terminal fragment of Venus (VC), thus generating a diploid collection. Then, each strain of the diploid collection expressing both the VN fusion and the VC fusion is analyzed by fluorescence microscopy, and the images are collected and quantitatively analyzed.











