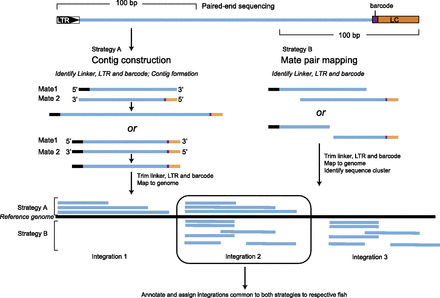
Strategies for mapping retroviral integrations. Paired-end sequencing was performed to capture the site of the retroviral integration (designated by LTR–retroviral 3′ long terminal repeat) and the linker cassette (LC) that contains the “barcode” identifier for the specific sample. Two strategies were used to map the integrations as this proved to be less error-prone than either strategy alone. In Strategy A, pairwise alignment of paired-end reads was performed to create contigs, and the resulting contigs were mapped to the zebrafish genome. Only contigs that mapped unambiguously were considered for identifying integrations. In Strategy B, each read from corresponding paired-ends was mapped independently, and colocalization in the correct orientation (pointing at each other) was used as the criterion for correct mapping. Integrations that mapped to the same genomic coordinates by both strategies were used for identification of integration events.











