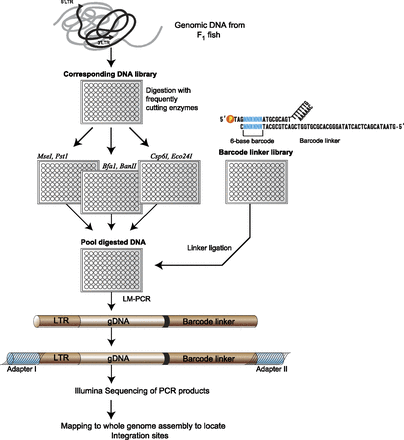
Overview of high-throughput strategy to identify retroviral integrations using a next-generation sequencing platform. Genomic DNAs corresponding to individual F1 fish were digested with three sets of restriction enzymes in parallel. After heat-inactivation of the restriction enzymes, the digested samples were then pooled together and ligated with DNA linkers, each containing a unique 6-bp barcode that indexes the F1 fish. The linker ligated DNA fragments were amplified by linker-mediated PCR using linker and viral LTR specific primers to amplify the adjacent genomic DNA sequences. The LTR/gDNA/linker amplicons are subsequently ligated to Illumina paired-end adapters and sequenced using the Illumina sequencing platform.











