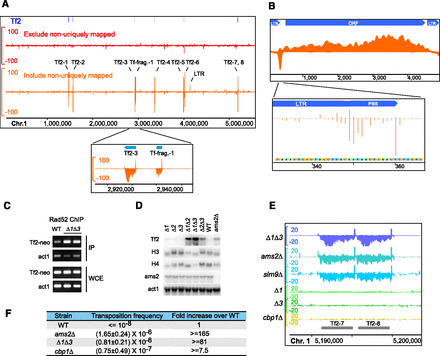
Rad52 enrichment on retrotransposon cDNA in histone dosage mutants. (A) Rad52 SPI-seq read distribution along chromosome 1 in Δ1Δ3. Enrichment at Tf2 retrotransposons was detected when nonuniquely aligned reads were randomly assigned and visualized. (B) Rad52 SPI-seq reads from Δ1Δ3 were aligned to a full-length Tf2 sequence (GenBank L10324.1). (Bottom panel) The unaveraged read numbers at primer-binding site (PBS). (C) ChIP-PCR detection of Rad52 enrichment on Tf2-12-neoAI. (D) Northern blotting analysis of Tf2 transcription in H3-H4 gene deletion mutants and ams2Δ. (E) Rad52 SPI-seq analysis of histone deletion mutants and mutants with higher-than-normal Tf2 expression. The tandem-linked Tf2-7 and Tf2-8 and their surrounding regions are shown. (F) Tf2 transposition frequency determined using Tf2-12-neoAI. The standard deviation was obtained from four independent cultures. The units on the y-axes of the browser views in A and E are reads per 10 million.











