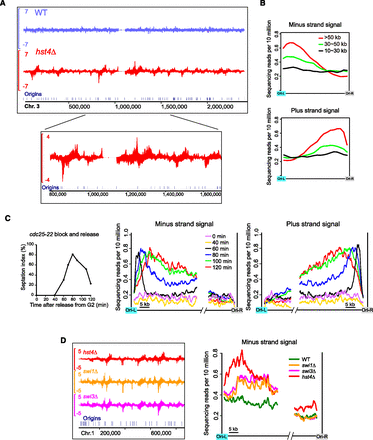
Replication-coupled Rad52 enrichment patterns in hst4Δ. (A) Rad52 SPI-seq read distribution along chromosome 3. (B) Averaged SPI-seq signal within inter-origin regions of different sizes. Inter-origin regions were classified into three groups according to their sizes. We divided each inter-origin region to 100 bins of equal sizes. The average read number within each bin was calculated, and a sliding window with a window size of 10 bins and a step size of five bins was applied to smooth the curves. (C) Rad52 SPI-seq analysis of hst4Δ cells synchronously released into cell cycle from a G2 arrest. The septation index (percentage of septated cells) shown on the left was measured with Calcofluor staining. The sequencing reads from inter-origin regions >50 kb were used to draw the average plots shown on the right. We first calculated the average read number at each nucleotide position within the first 35 kb and the last 15 kb of the inter-origin regions, and then applied a sliding window with a window size of 1 kb and a step size of 0.1 kb to smooth the curves. (D) Rad52 SPI-seq analysis of swi1Δ and swi3Δ. Average plots were drawn as in C. In A and D, a bandwidth of 500 bp was used for kernel density estimation, and the units on the y-axes of the browser views are reads per 10 million.











