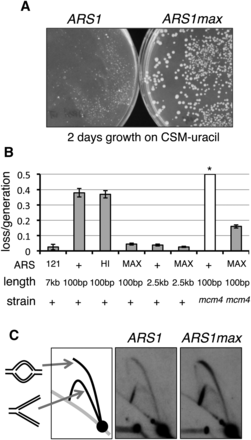
Optimized synthetic ARS function. (A) ARS1 and ARS1max sequences were cloned into a URA3 vector. Yeast were transformed with resultant plasmids and grown for 2 d on medium lacking uracil. Colony size is representative of ARS function. (B) Plasmid loss assays were performed on CEN vectors containing ARSs indicated: (+) ARS1; (HI) ARS1hi1; (MAX) ARS1max1.1; (121) ARS121. A control vector YCP121, bearing a 7-kb ARS121 fragment is shown (121) as an example of very strong ARS function. (*) The loss rate of wild-type ARS1 in mcm4-chaos3 cells (mcm4) was too high to be measured. (C) Genomic DNA replication was assayed in strains with ARS1 and ARS1max integrated into the native chromosomal locus. The signal ratio of the top (bubble) arcs to the bottom (Y) arcs is indicative of replication initiation at the assayed locus.











