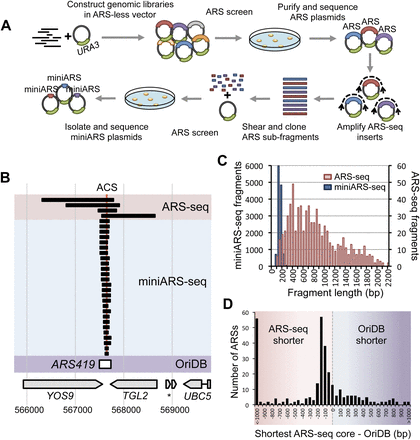
High-throughput, high-resolution mapping of ARSs using ARS-seq and miniARS-seq. (A) Genomic libraries were constructed in a URA3 vector lacking an ARS. Yeast were transformed with these libraries for selection of ARS-containing plasmids. ARS plasmids were isolated from pooled yeast colonies and sequenced using insert-flanking primers (ARS-seq, top row). Inserts from the ARS-seq plasmid pools were amplified using vector-specific primers. Randomly sheared and size-selected fragments were cloned into an ARS-less vector and rescreened for ARS function (miniARS-seq, bottom row). (B) A sample locus (at ARS419) comparing results of ARS-seq (red highlight), miniARS-seq (blue highlight), and OriDB annotation (purple highlight). The best match of the ACS motif is indicated at the top (red vertical line). Corresponding coordinates on chromosome 4 and annotated nearby genes are shown at the bottom. (C) Size distributions of ARS-seq and miniARS-seq fragments listed in Supplemental Tables 1 and 2. (D) Distribution of the size differences between OriDB-annotated confirmed ARSs and the corresponding shortest ARS-seq/miniARS-seq fragments or inferred functional cores (see Methods).











