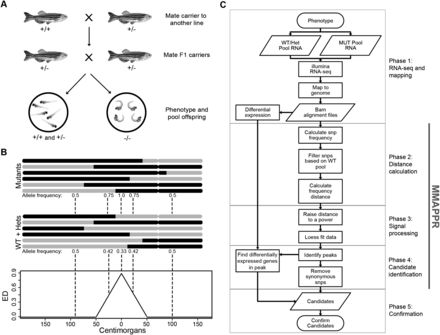
Schematic overview of MMAPPR. (A) Mating scheme. Fish from the mutant line are outcrossed and then F1 progeny, heterozygous for the mutation, are crossed. Pools of phenotypically mutant and phenotypically wild-type fish are then sorted. (B) Schematic representation of allelic segregation between wild-type and mutant pools. (Black) Genomic regions inherited from the mutant carrier line; (gray) regions inherited from the outcross line. (Bottom panel) A plot of the expected Euclidean distances calculated from the allele frequencies in the two pools. (C) Flowchart of the analysis steps incorporated in the MMAPPR algorithm. Each phase of the pipeline is shown by gray boxes, and the portion of the pipeline processed by the MMAPPR software package is shown by a vertical gray bar.











