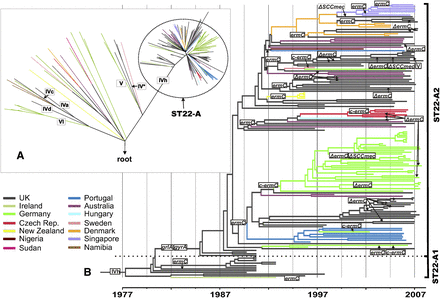
Phylogeny of ST22 and the emergence of MRSA clones. (A) Maximum likelihood phylogenetic tree of ST22 isolates. The tree was rooted by using the distantly related S. aureus isolate MSSA476 as an outgroup. Colors indicate the isolates' countries of origin. Roman numerals indicate acquisitions of structurally different SCCmec elements, which cause methicillin resistance. (B) Maximum clade credibility tree of the ST22-A clade based on BEAST analysis using a variable clock rate (uncorrelated lognormal) model. Tips of the tree are constrained by isolation dates, the time scale is shown at the bottom. Gains and losses (Δ) of genetic determinants for resistance to methicillin (SCCmec IVh), fluoroquinolones (point mutations in grlA and gyrA), erythromycin (plasmid-encoded ermC), and clindamycin (mutations in ermC leader peptide region, c-ermC) have been mapped on the tree by applying the parsimony criterion.











