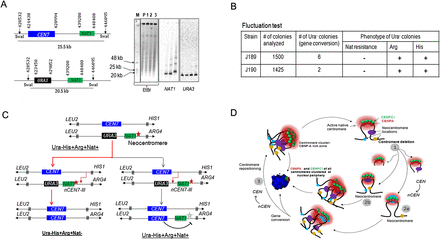
Centromere repositioning occurs due to inactivation of neocentromeres by gene conversion. (A) Schematics showing a region across CEN7 and expected size of the SwaI fragment released either from the unaltered (25.5 kb) or CEN7-deleted (20.5 kb) homolog of Chr7 (left panel). SwaI-digested genomic DNA plugs from the parent (RM1000AH-cen7-6.5kbΔ), and three NAT1 integrants were separated on a CHEF gel (EtBr stained CHEF gel is shown here). Probing with NAT1 or URA3 sequences confirmed integration of NAT1 into CEN7-deleted neocentric Chr7 in these two integrants (J189 and J190). (B) A table summarizes the phenotypes of the markers of the segregants. Fluctuation test analysis in two independent NAT1 integrants (J189 and J190) of J181 (RM1000AH-cen7-6.5kbΔ) revealed the loss of NAT1 along with URA3. This result suggests that the neocentromere has been replaced along with URA3 by gene conversion. (C) Schematic showing possible fates of the segregants after gene conversion depending on the site of recombination initiation. (Red line) The observed event. (Black line) An alternative event. (Red star) Active neocentromere; (gray star) inactive neocentromere. (D) A possible mechanism of the maintenance of centromere function at the native physical chromosomal location. Similar to budding yeast S. cerevisiae, centromeres are always clustered toward the periphery of a nucleus throughout the cell cycle in C. albicans. A 3D reconstructed image showing peripheral localization of the kinetochore cluster immunostained with anti-CENPA/Cse4 (red) and anti-CENPC1/Mif2 (green) antibodies in a nucleus (blue, DAPI) in C. albicans. Clustering of kinetochores/centromeres is crucial for the integrity of the kinetochore ensemble as well as the fidelity of chromosome segregation in both S. cerevisiae and C. albicans (Anderson et al. 2009). We propose that clustering of all native centromeres into a specific 3D scaffold at the nuclear periphery creates a zone of high CENPA (and other kinetochore proteins including CENPC1) local concentration (pink). Potential neocentromere sites exist at centromere proximal regions (oval shapes of blue, brown, and violet colors) at least partly, due to their physical proximity to the CENPA-rich zone. Under normal conditions, the native centromere occupies the most preferred position in the centromere cluster and keeps adjacent potential neocentromere sites away from the CENPA-rich zone by providing steric hindrance. Removal of centromeric regions of various lengths brings centromere proximal regions in close proximity to the CENPA-rich zone (1). The new region now becomes a part of the CENPA-rich nuclear domain and assembles CENPA chromatin to form a neocentromere (2a and 2b). Gene conversion can inactivate an established neocentromere by replacing it with the homologous sequence that lacks the epigenetic information for CENPA assembly (3). This process is accompanied by acquisition of the native centromere from a homologous chromosome leading to repositioning of the centromere back to its original location.











