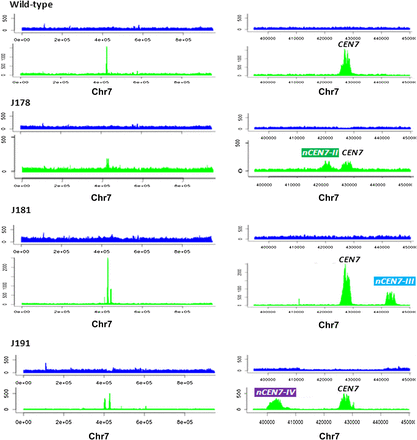
ChIP-seq analyses revealed that neocentromeres are always and exclusively formed at a close proximity to the native CEN. Relative number of sequence reads from the whole cell lysate (upper panels) or CENPA-Prot A or MycMif2/CENPC1 ChIP DNA (lower panels) in wild-type strain J200, Cen7Δ-deleted strains J178 (RM1000AH-cen7Δ), J181 (RM1000AH-cen7-6.5kbΔ), or J191 (RM1000AH-cen7-30kbΔ) mapped neocentromeres (nCENs) within 15 kb from the deleted region. (Left panels) Distribution of sequence reads across the entire chr7; (right panels) zoomed-in view of neocentromeric location with respect to the native CEN7 in each case. Chromosome coordinates are shown in the x-axis.











