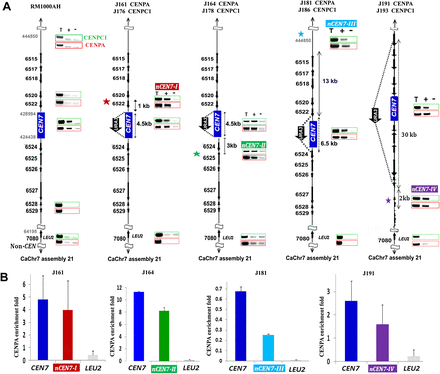
Neocentromeres, like centromeres, preferentially form on specific chromosomal locations in a nonrandom fashion independent of the length of the deleted region in C. albicans. (A) CENPA and CENPC1 ChIP analyses in RM1000AH-cen7Δ, RM1000AH-cen7-6.5 kbΔ, or RM1000AH-cen7-30kbΔ clones mapped neocentromeres (nCENs) within 15 kb from the deleted region. C. albicans is a diploid yeast and carries two homologs of each chromosome. Only one homolog of Chr7 where CEN7 has been replaced by URA3 is shown. CENPA enrichment shown at the native CEN7 location is contributed by an unaltered homolog and is shown as a positive control for CENPA ChIP assays. Thick black arrows along with the Orf numbers show the gene arrangement and the direction of transcription. One representative clone from each class (see Supplemental Figs. S3C, S4B for other clones) is shown. Class I neocentromeres (nCEN7-I) formed on Orf19.6520 and Orf19.6522 (10/11 RM1000AH-cen7Δ and 4/5 RM1000AH-cen-6.5kbΔ transformants), Class II neocentromeres (nCEN7-II) mapped to Orf19.6526 and Orf19.6525 (1/11 RM1000AH-cen7Δ), Class III neocentromeres (nCEN7-III) mapped on an Orf-free region—Assembly 21, CaChr7 Coordinates 442000–445000 (1/5 RM1000AH-cen-6.5kbΔ transformant)—and nCEN7-IV mapped on Orf19.6531, Orf19.6532, Orf19.6533, Orf19.6534 (2/2 RM1000AH-cen7-30kbΔ transformants). Parent RM1000AH did not show CENPA or CENPC1 enrichment on these neocentromere sites. Coordinates of the control non-CEN (LEU2) region on Chr7 in Assembly 21 are 64195–64440 (Orf 19.7080). (T) Total DNA; (+) with Ab; (−) without Ab (beads only). CENPA ChIP profiles are shown in red boxes and the same for CENPC1 are shown in green boxes. Enrichment of CENPA at the URA3 location was not shown here but presented as a separate figure (Supplemental Fig. S5A). (B) CENPA ChIP qPCR analysis showing fold enrichment of CENPA on native and neocentromeres.











