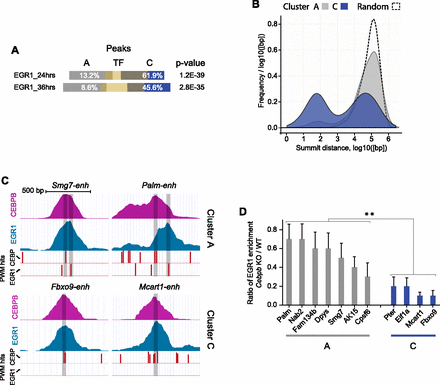
Two modes of EGR1 cobinding at A and C cluster regions. (A) Bar diagram displaying overlap between peaks defined by EGR1 binding (24 h or 36 h, yellow bars) and CEBP binding. A or C cluster CEBP peak overlaps are shown (white numbers indicate proportions) with P-values of hypergeometric tests (Fisher's one-tailed) of similarity. (B) Diagram showing the distribution of distances between CEBP peak summits and the closest EGR1 (36-h) peak. A (gray) and C (blue) cluster peaks and a randomly distributed set (dashed line) are included. (C) CEBP and EGR1 ChIP read coverage and PWM hit tracks from representative A (top panels) and C (bottom panels) cluster regions. Vertical red bars indicate CEBP and EGR1 PWM hits; height of bars identity score; and transparent black bars show position of peak summits. (D) EGR1-ChIP using livers of mice deficient for Cebpb and wild-type livers. (Y-axis) Ratio of enrichment for Cebpb-null versus wild-type experiments. EGR1 bound regions overlapping with CEPB A cluster (gray) and C cluster (blue) peaks are shown. N = 3–4. Error bars, SEM. (**) P < 0.01, Mann-Whitney test, two-tailed. For gene names, see Supplemental Table S10.











