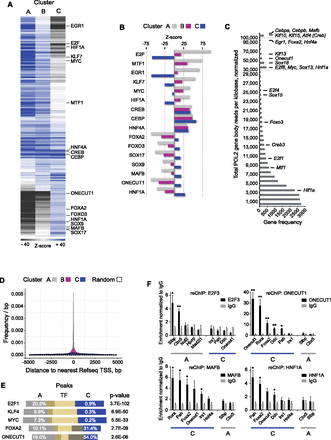
The CEBP temporal binding patterns display two sets of cis-regulatory code. (A) Representation of motif frequencies in each temporal cluster. Hierarchical clustering based on Z-scores of 210 position weight matrices (PWMs). Each row indicates a specific motif (PWM), while columns represent each binding cluster (A, B, or C). Color scale indicates representation relative to background set (overrepresented, dark blue; underrepresented, dark gray; no difference, white). Z-score scale is cut off at ±40 for clarity. Example PWMs are shown in black text. (B) Selected transcription factors with clear binding sequence overrepresentation in A and B clusters (upper six), all clusters (middle three), or only the C cluster (lower seven). Clusters are denoted by color. (C) Expression level frequency distribution (POL2 gene body read coverage) of all genes (reads per kilobase, sum of eight time points, normalized, above 0.5) with candidate transcription factors indicated. (D) Distances from A, B, and C cluster peak summits or random positions to the most proximal RefSeq transcription start site (TSS). (E) External ChIP-seq data peak regions (yellow bars) showing overlaps with CEBP A and C cluster regions (white numbers indicate proportions) and P-value of hypergeometric test (Fisher's one-tailed) of overlap similarity. (F) Sequential ChIP (reChIP) assessing co-occupancy at A and C cluster regions. Anti-CEBP was used as first-round antibody (recognizing both CEBPA and CEBPB), with second-round IgG or antibody against indicated TFs. Enrichments are normalized to IgG levels. N = 2–5. Error bars, SEM. (*) P < 0.05; (**) P < 0.01, t-test versus IgG enrichments. For gene names, see Supplemental Table S10.











